Ordered by weight (if set) and creation date.
Research
Research in the Stevens lab is concerned with the process of protein sorting and membrane assembly in yeast cells. Using yeast molecular genetics, we have identified a large number of genes required for the correct targeting and transport of proteins to the membrane-bounded organelle called the vacuole. These vacuolar protein sorting (VPS) genes have been found to encode proteins such as a dynamin-like GTPase, a protein-sorting receptor, a protein kinase, a lipid kinase, a RAS inhibitor-like protein, and an increasingly large number of proteins involved in transport vesicle targeting/fusion such as Rab-like GTPases and SNARE proteins. To characterize the function of some of these proteins we use biochemical, cell biological and molecular genetic approaches. Biochemical approaches are being used to isolate a number of the VPS proteins and to study the membrane-associated protein complexes in which they are found.
Figure 1
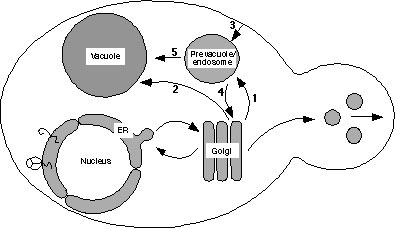
Schematic drawing of the yeast endomembrane pathways. Proteins exiting the Golgi complex can be transported by either the CPY pathway (arrow 1) through the prevacuole to the vacuole, or by the ALP pathway (arrow 2). Proteins can also reach the prevacuole by endocytosis (arrow 3), and proteins in the prevacuole transit a common pathway to the vacuole (arrow 5). Golgi membrane proteins such as DPAP A are retrieved from the prevacuole back to the Golgi (arrow 4).
The group also has a long-standing interest in the assembly, targeting, structure and function of the vacuolar H+-translocating ATPase (V-ATPase; see figure). The V-ATPase complex consists of fourteen subunits, and all but one of these are encoded by a single yeast gene. The large hydrophobic V-ATPase “a” subunit has two isoforms, Vph1 and Stv1, with the Vph1-associated V-ATPase complex localizing to the vacuole membrane and the Stv1-associated V-ATPase restricted to Golgi and endosomal membranes. The mechanism of differential localization of these two forms of the yeast V-ATPase is under active investigation in the lab. We are investigating the proteins responsible for maintaining this differential localization, as well as the protein-based signals that specify the distinct localizations.
We have also identified five genes that encode proteins required for V-ATPase complex assembly but are not themselves part of the final V-ATPase enzyme complex. These five proteins reside in the yeast cell endoplasmic reticulum and constitute the dedicated assembly machinery for the V-ATPase. A number of molecular genetic and biochemical approaches are being taken to characterize the assembly complex and to study the interaction of this assembly complex with V-ATPase subunits along the assembly pathway within the endoplasmic reticulum.
Figure 2
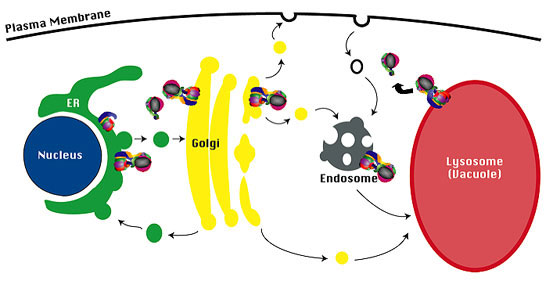
Targeting of V-ATPase complexes.The membrane sector of the V-ATPase complex (V 0 sector) is assembled in the ER, and the assembled V-ATPase complex is then transported to the Golgi complex. The Stv1-containing V-ATPase complex cycles between the endosome and the Golgi complex, and the Vph1-containing V-ATPase complex is transported to the limiting vacuole membrane.
The lab has also employed ancestral gene reconstruction to investigate the V-ATPase enzyme complex in more detail. Ancestral reconstruction of the two V-ATPase subunit a isoforms has generated the most likely predecessor gene prior to gene duplication. This Anc.a protein functions with the remaining 13 V-ATPase subunits and has characteristics of both the Stv1- and Vph1-containing V-ATPase complexes. Ancestral reconstruction of the most likely predecessor of the Vma3 and Vma11 proteolipid V-ATPase subunits has led to the synthesis of an Anc.3-11 the functions with the remaining 12 V-ATPase subunits to form a functional enzyme complex. The Anc.3-11 protein allows the assembly of a proteolipid ring with two different polypeptides (Anc.3-11 and Vma16) rather than the modern-day ring with three different polypeptides (Vma3, Vma11 & Vma16). These investigations have revealed important insights into the modern-day V-ATPase complex as well as the evolution of this V-ATPase molecular machine.
Figure 3
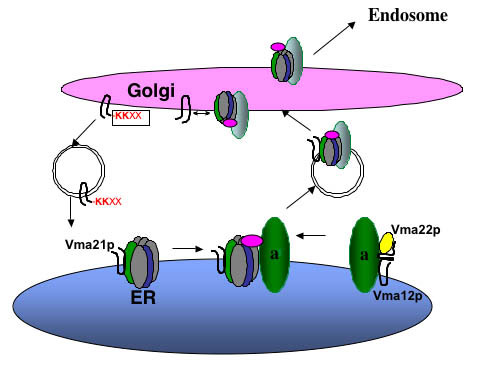
Assembly of the Vo sector in the ER. Vma21p interacts with the proteolipid ring V-ATPase subunits of Vma3p (c), Vma11p (c’), and Vma16p (c’’). Vma12p and Vma22p form a complex on the ER membrane and interact with newly synthesized “a” subunit (Vph1p and Stv1p), and then Vma21p “escorts” this Vo sector into vesicles budding from the ER membrane. Vma21p accompanies the V-ATPase to the Golgi complex, where its C-terminal KKXX ER retention signal is recognized by the COPI machinery and this assembly/escort factor is recycled to the ER. The assembled V-ATPase is then transported on to the endosome and/or vacuole.
| Attachment | Size |
|---|---|
| stevens_fig1.jpg | 19.62 KB |
| stevens_fig2.jpg | 31.3 KB |
| stevens_fig3.jpg | 26.01 KB |
